About 3DIV
3D genome organization is tightly coupled with gene regulation in various biological processes and diseases. 3D Interaction Viewer and Database (3DIV) is a database providing chromatin interaction visualization in a variety of options from one-to-all chromatin interaction with epigenetic annotation to unique dynamic browsing tools allowing examination of large-scale genomic rearrangement mediated impacts in cancer 3D genome. 3DIV will be the most comprehensive resource to explore gene regulatory effects of both normal and cancer 3D genome.
Hi-C
3DIV provides querying list of significant interacting partner locus, visualization, and comparative analysis of 3D chromatin interaction across about 400 samples.
Capture Hi-C
3DIV provides promoter capture Hi-C (pcHi-C) results across 28 normal human cell/tissue types, a great resource in identifying target genes of disease-associated genetic variants.
Cancer Hi-C
3DIV provides unique visualization and manipulation tools that allows user to generate rearranged 3D genome by selecting listed SVs, creating own SVs, and providing order of rearranged chromosomes.
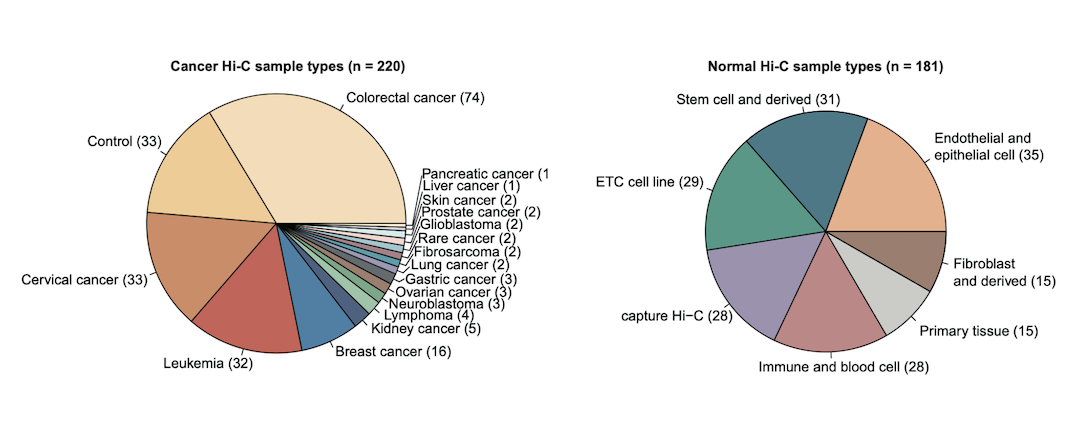
Samples
Cancer types
Colon cancer specific Structural Variations
References
-
3DIV Update for 2021: a comprehensive resource of 3D genome and 3D cancer genome
Kyukwang Kim, Insu Jang, Mooyoung Kim, Jinhyuk Choi, Min-Seo Kim, Byungwook Lee, and Inkyung Jung
Nucleic Acids Res. 2021 Jan; Vol 49, Issue D1 (Database issue):D38–D46.
-
3DIV: A 3D-genome Interaction Viewer and databaseWeb site
Dongchan Yang, Insu Jang, Jinhyuk Choi, Min-Seo Kim, Andrew J Lee, Hyunwoong Kim, Junghyun Eom, Dongsup Kim, Inkyung Jung and Byungwook Lee
Nucleic Acids Res. 2018 Jan; Vol 46, Issue D1 (Database issue):D52-57.
-
A compendium of promoter-centered long-range chromatin interactions in the human genome
Inkyung Jung, Anthony Schmitt, Yarui Diao, Andrew J. Lee, Tristin Liu, Dongchan Yang, Catherine Tan, Junghyun Eom, Marilynn Chan, Sora Chee, Zachary Chiang, Changyoun Kim, Eliezer Masliah, Cathy L. Barr, Bin Li, Samantha Kuan, Dongsup Kim & Bing Ren
Nature Genetics volume 51, pages 1442–1449(2019).